The July and August meetings of the New England R Users group focused on two different aspects of R performance: parallel processing techniques and the effects of compiler & library selection when compiling the R executable itself.
It was during Amy Szczepanski’s excellent introduction to multicore, Rmpi, and foreach (slides here) that she mentioned that the nice people who compile R for the Mac use optimized libraries to improve its performance. Amy works at the University of Tennessee’s Center for Remote Data Analysis and Visualization where they build and run machines with tens of thousands of cores, so her endorsement carries a lot of weight. I think Amy mentioned that she had benchmarked the open source vs. Revolutions distributions on her Mac, but I can’t find it in her slides and, well, in one ear and out the other….
It was the comprehensive presentation by IBM’s Vipin Sachdeva (slides here) showing 15-20X speedups through compiler and library selection that made me want to try a couple of benchmarks myself. And my recent it’s-about-time upgrade to 2.11 seemed like the perfect opportunity.
Open source vs. Revolutions Community R
Performance is one of the advantages claimed by Revolution Analytics for its distributions, with their product page promising “optimized libraries and compiler techniques run most computation-intensive programs significantly faster than Base R” even with their free, Community edition. I have heard good things about its performance on Windows, so I was curious to see if it provides an improvement over the already-optimized Mac binary.
Benchmarking Methodology (or lack thereof)
First, some disclaimers: I am not a serious benchmarker and have made no special effort for statistical rigor. I am just looking for order-of-magnitudes here, so I kept a normal number of programs running in the background, like Firefox and OpenOffice, though nothing was doing anything substantial and I avoided any user input while each test ran. My machine is the short-lived, late-2008, aluminum unibody 13″ MacBook (MacBook5,1) with 4GB RAM and Mac OS X Leopard 10.5.8 running the 32-bit kernel. It has a 2.4GHz Core 2 Duo — nothing special.
For my tests, I ran the standard R Benchmark 2.5 available from AT&T ‘s benchmarking page which performs various matrix and vector calculations — perfect for discerning the effects of such optimized libraries. I kept the defaults, such as running each test 3 times, and installed the required “SuppDists” package. I tested the open source 2.10.1 32-bit version I already had on my machine and then installed Revolution’s 2.10.1-based 64-bit community edition. I should have repeated the test with the open source 64-bit edition, but I didn’t think of it at the time (I told you I wasn’t serious about this), so instead I later re-ran the benchmark with the 32- and 64-bit versions of the open source 2.11.1 to check if there are any significant 32-vs-64 differences.
Results
It didn’t take long to realize that the Revolutions community edition was not going to fare well. During just the fourth benchmark, 2800×2800 cross-product matrix (b = a’ * a), there was a pregnant pause in the output while my laptop’s fans kicked in and soon spun up to full force. It took nearly 25 seconds to complete each turn of that one test where the open source 2.10.1 had finished in less than one tenth the time. (The complete output for each test is at the end of this post.)
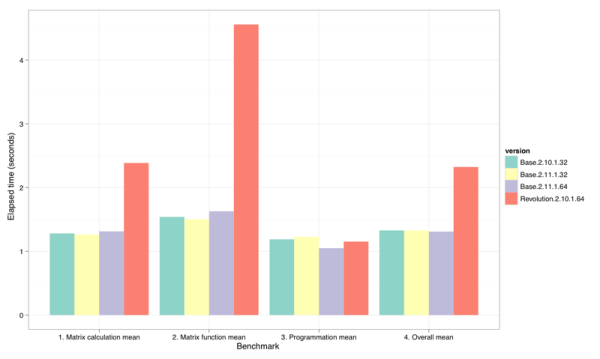
Figure 1: Summary-level benchmark results. (Smaller bars are better.)
Figure 1 shows the geometric means of the elapsed times for each benchmark section as reported by R Benchmark 2.5. Clearly the Revolutions distribution did significantly worse on the matrix benchmarks. Figure 2 drills into the individual benchmarks to show the roughly 2-8X difference on the five slowest matrix benchmarks. Only on the sixth, Grand common divisors of 400,000 pairs (recursion), was the slowdown matched by the base 64-bit distribution. Only on Revolution’s fastest benchmark,2400×2400 normal distributed random matrix ^1000 all the way at the bottom of Figure 2, did the 64-bit versions hold a distinct (and roughly equal) advantage over their 32-bit brethren.

Figure 2: Individual benchmark results. (Smaller bars are better.)
vecLib: BLAS, LAPACK, and built into the Mac
So, no surprise — Amy was right. The off-the-shelf open source distributions of R for the Mac are already optimized. But how? Vipin walked us through all the different choices for BLAS and LAPACK libraries, not to mention the different C and FORTRAN compilers and their optimization flags. How can we know what’s being used by a given distribution? Well, it turns out that R makes it easy to find out with the config options to “R CMD”:
$ R CMD config BLAS_LIBS-L/Library/Frameworks/R.framework/Resources/lib/i386 -lRblas
$ ls -l /Library/Frameworks/R.framework/Resources/lib/i386/libRblas.dylib
lrwxr-xr-x 1 root admin 17 Oct 7 00:58 /Library/Frameworks/R.framework/Resources/lib/i386/libRblas.dylib -> ../libRblas.dylib
$ ls -l /Library/Frameworks/R.framework/Resources/lib/libRblas.dylib
lrwxr-xr-x 1 root admin 21 Oct 7 00:58 /Library/Frameworks/R.framework/Resources/lib/libRblas.dylib -> libRblas.vecLib.dylib
Following the symbolic link, “libRblas.vecLib.dylib” is the library being used for BLAS. A quick consultation with Google reveals that “vecLib” is Apple’s vector library from their “Acceleration” framework in Mac OS. Here’s what Apple’s Developer site says about vecLib’s BLAS and LAPACK components::
Basic Linear Algebra Subprograms (BLAS)
VecLib also contains Basic Linear Algebra Subprograms (BLAS) that use AltiVec technology for their implementations. The functions are grouped into three categories (called levels), as follows:
- Vector-scalar linear algebra subprograms
- Matrix-vector linear algebra subprograms
- Matrix operations
A Readme file is included that contains the following sections:
- Descriptions of functions
- Comparison with BLAS (Basic Linear Algebra Subroutines)
- Test methodology
- Future releases
- Compiler version
LAPACK
LAPACK provides routines for solving systems of simultaneous linear equations, least-squares solutions of linear systems of equations, eigenvalue problems, and singular value problems. The associated matrix factorizations (LU, Cholesky, QR, SVD, Schur, generalized Schur) are also provided, as are related computations such as reordering of the Schur factorizations and estimating condition numbers. Dense and banded matrices are handled, but not general sparse matrices. In all areas, similar functionality is provided for real and complex matrices, in both single and double precision.
Also, see <http://netlib.org/lapack/index.html.
As Vipin had demonstrated, using a fast BLAS and LAPACK libraries can make all the difference in the world (well, 20X or so). And since Apple controls the horizontal and vertical on their platform, it shouldn’t be a surprise that vecLib is fast on their hardware and OS. The real question is why doesn’t Revolutions simply link to vecLib too? It can’t be because their libraries are better (they clearly aren’t). Nor could they be afraid of competing with their Enterprise edition because, according to this edition comparison chart, they don’t offer an enterprise edition for the Mac. Perhaps they’re simply not that familiar with the platform and don’t know about vecLib. I know I didn’t know anything about it until these tests prompted me to consult The Google.
Your mileage may vary
Google also pointed me to this recent discussion on the R-SIG-MAC mailing list in which Simon Urbanek refers to a serious bug in vecLib which prevents it from spawning threads and shows timings from a “tcrossprod” benchmark on a new 2.66GHz Mac Pro:
vecLib 6.43
ATLAS serial 4.80
MKL serial 4.30
MKL parallel 0.90
ATLAS parallel 0.71
So there still seems to be plenty of opportunity to beat vecLib if you’re willing to compile R and mix and match BLAS libraries. For the rest of us, the open source distribution offers the best bang for the buck.
Maybe I need to ask Vipin to take a look at my Mac at the next meeting….
Complete Output
open source 2.10.1 32-bit
R version 2.10.1 (2009-12-14)
Copyright (C) 2009 The R Foundation for Statistical Computing
ISBN 3-900051-07-0
R is free software and comes with ABSOLUTELY NO WARRANTY.
You are welcome to redistribute it under certain conditions.
Type 'license()' or 'licence()' for distribution details.
Natural language support but running in an English locale
R is a collaborative project with many contributors.
Type 'contributors()' for more information and
'citation()' on how to cite R or R packages in publications.
Type 'demo()' for some demos, 'help()' for on-line help, or
'help.start()' for an HTML browser interface to help.
Type 'q()' to quit R.
[R.app GUI 1.31 (5537) i386-apple-darwin8.11.1]
[Workspace restored from /Users/jbreen/.RData]
> source("/Users/jbreen/Desktop/R-benchmark-25.R")
Loading required package: Matrix
Loading required package: lattice
Loading required package: SuppDists
R Benchmark 2.5
===============
Number of times each test is run__________________________: 3
I. Matrix calculation
---------------------
Creation, transp., deformation of a 2500x2500 matrix (sec): 1.16766666666666
2400x2400 normal distributed random matrix ^1000____ (sec): 1.14666666666667
Sorting of 7,000,000 random values__________________ (sec): 1.26266666666667
2800x2800 cross-product matrix (b = a' * a)_________ (sec): 2.461
Linear regr. over a 3000x3000 matrix (c = a \ b')___ (sec): 1.42233333333333
--------------------------------------------
Trimmed geom. mean (2 extremes eliminated): 1.27997944504873
II. Matrix functions
--------------------
FFT over 2,400,000 random values____________________ (sec): 1.333
Eigenvalues of a 640x640 random matrix______________ (sec): 1.23633333333333
Determinant of a 2500x2500 random matrix____________ (sec): 1.65233333333333
Cholesky decomposition of a 3000x3000 matrix________ (sec): 1.712
Inverse of a 1600x1600 random matrix________________ (sec): 1.65633333333334
--------------------------------------------
Trimmed geom. mean (2 extremes eliminated): 1.53942494113224
III. Programmation
------------------
3,500,000 Fibonacci numbers calculation (vector calc)(sec): 1.21066666666667
Creation of a 3000x3000 Hilbert matrix (matrix calc) (sec): 0.919333333333332
Grand common divisors of 400,000 pairs (recursion)__ (sec): 1.252
Creation of a 500x500 Toeplitz matrix (loops)_______ (sec): 1.38000000000001
Escoufier's method on a 45x45 matrix (mixed)________ (sec): 1.105
--------------------------------------------
Trimmed geom. mean (2 extremes eliminated): 1.18758233972606
Total time for all 15 tests_________________________ (sec): 20.9173333333333
Overall mean (sum of I, II and III trimmed means/3)_ (sec): 1.32762395973868
--- End of test ---
Revolution R community 2.10.1 64 bit
R version 2.10.1 (2009-12-14)
Copyright (C) 2009 The R Foundation for Statistical Computing
ISBN 3-900051-07-0
R is free software and comes with ABSOLUTELY NO WARRANTY.
You are welcome to redistribute it under certain conditions.
Type 'license()' or 'licence()' for distribution details.
R is a collaborative project with many contributors.
Type 'contributors()' for more information and
'citation()' on how to cite R or R packages in publications.
Type 'demo()' for some demos, 'help()' for on-line help, or
'help.start()' for an HTML browser interface to help.
Type 'q()' to quit R.
REvolution R version 3.2: an enhanced distribution of R
REvolution Computing packages Copyright (C) 2010 REvolution Computing, Inc.
Type 'revo()' to visit www.revolution-computing.com for the latest
REvolution R news, 'forum()' for the community forum, or 'readme()'
for release notes.
[R.app GUI 1.30 (5511) x86_64-apple-darwin9.8.0]
[Workspace restored from /Users/jbreen/.RData]
trying URL 'http://watson.nci.nih.gov/cran_mirror/src/contrib/SuppDists_1.1-8.tar.gz'
Content type 'application/x-gzip' length 139864 bytes (136 Kb)
opened URL
==================================================
downloaded 136 Kb
* installing *source* package ‘SuppDists’ ...
** libs
** arch - x86_64
g++ -arch x86_64 -I/opt/REvolution/Revo-3.2/Revo64/R.framework/Resources/include -I/opt/REvolution/Revo-3.2/Revo64/R.framework/Resources/include/x86_64 -I/usr/local/include -fPIC -g -O2 -c dists.cc -o dists.o
g++ -arch x86_64 -dynamiclib -Wl,-headerpad_max_install_names -undefined dynamic_lookup -single_module -multiply_defined suppress -L/usr/local/lib -o SuppDists.so dists.o -F/opt/REvolution/Revo-3.2/Revo64/R.framework/.. -framework R -Wl,-framework -Wl,CoreFoundation
** R
** preparing package for lazy loading
** help
*** installing help indices
** building package indices ...
* DONE (SuppDists)
The downloaded packages are in
‘/private/var/folders/+s/+snLnQILHz4Kn8jMFYEERE++-4+/-Tmp-/RtmpVqNN4f/downloaded_packages’
> source("/Users/jbreen/Desktop/R-benchmark-25.R")
Loading required package: Matrix
Loading required package: lattice
Loading required package: SuppDists
R Benchmark 2.5
===============
Number of times each test is run__________________________: 3
I. Matrix calculation
---------------------
Creation, transp., deformation of a 2500x2500 matrix (sec): 1.18633333333333
2400x2400 normal distributed random matrix ^1000____ (sec): 0.463333333333334
Sorting of 7,000,000 random values__________________ (sec): 1.10266666666667
2800x2800 cross-product matrix (b = a' * a)_________ (sec): 24.786
Linear regr. over a 3000x3000 matrix (c = a \ b')___ (sec): 10.3813333333333
--------------------------------------------
Trimmed geom. mean (2 extremes eliminated): 2.38580368558367
II. Matrix functions
--------------------
FFT over 2,400,000 random values____________________ (sec): 1.32633333333335
Eigenvalues of a 640x640 random matrix______________ (sec): 1.631
Determinant of a 2500x2500 random matrix____________ (sec): 8.66366666666666
Cholesky decomposition of a 3000x3000 matrix________ (sec): 6.70300000000001
Inverse of a 1600x1600 random matrix________________ (sec): 8.71566666666664
--------------------------------------------
Trimmed geom. mean (2 extremes eliminated): 4.55835668372464
III. Programmation
------------------
3,500,000 Fibonacci numbers calculation (vector calc)(sec): 1.10766666666666
Creation of a 3000x3000 Hilbert matrix (matrix calc) (sec): 0.818666666666672
Grand common divisors of 400,000 pairs (recursion)__ (sec): 3.529
Creation of a 500x500 Toeplitz matrix (loops)_______ (sec): 1.28933333333335
Escoufier's method on a 45x45 matrix (mixed)________ (sec): 1.072
--------------------------------------------
Trimmed geom. mean (2 extremes eliminated): 1.15254093929808
Total time for all 15 tests_________________________ (sec): 72.776
Overall mean (sum of I, II and III trimmed means/3)_ (sec): 2.32291395838888
--- End of test ---
open source 2.11.1 64-bit
R version 2.11.1 (2010-05-31)
Copyright (C) 2010 The R Foundation for Statistical Computing
ISBN 3-900051-07-0
R is free software and comes with ABSOLUTELY NO WARRANTY.
You are welcome to redistribute it under certain conditions.
Type 'license()' or 'licence()' for distribution details.
Natural language support but running in an English locale
R is a collaborative project with many contributors.
Type 'contributors()' for more information and
'citation()' on how to cite R or R packages in publications.
Type 'demo()' for some demos, 'help()' for on-line help, or
'help.start()' for an HTML browser interface to help.
Type 'q()' to quit R.
[R.app GUI 1.34 (5589) x86_64-apple-darwin9.8.0]
[Workspace restored from /Users/jbreen/.RData]
> source("/Users/jbreen/Desktop/R-benchmark-25.R")
Loading required package: Matrix
Loading required package: lattice
Attaching package: 'Matrix'
The following object(s) are masked from 'package:base':
det
Loading required package: SuppDists
Error in eval.with.vis(expr, envir, enclos) : object 'rMWC1019' not found
In addition: Warning message:
In library(package, lib.loc = lib.loc, character.only = TRUE, logical.return = TRUE, :
there is no package called 'SuppDists'
trying URL 'http://watson.nci.nih.gov/cran_mirror/bin/macosx/leopard/contrib/2.11/SuppDists_1.1-8.tgz'
Content type 'application/x-gzip' length 412485 bytes (402 Kb)
opened URL
==================================================
downloaded 402 Kb
The downloaded packages are in
/var/folders/+s/+snLnQILHz4Kn8jMFYEERE++-4+/-Tmp-//RtmpoWgMNZ/downloaded_packages
> source("/Users/jbreen/Desktop/R-benchmark-25.R")
Loading required package: SuppDists
R Benchmark 2.5
===============
Number of times each test is run__________________________: 3
I. Matrix calculation
---------------------
Creation, transp., deformation of a 2500x2500 matrix (sec): 1.193
2400x2400 normal distributed random matrix ^1000____ (sec): 0.455333333333336
Sorting of 7,000,000 random values__________________ (sec): 1.06133333333333
2800x2800 cross-product matrix (b = a' * a)_________ (sec): 2.70466666666667
Linear regr. over a 3000x3000 matrix (c = a \ b')___ (sec): 1.78500000000001
--------------------------------------------
Trimmed geom. mean (2 extremes eliminated): 1.3123313069964
II. Matrix functions
--------------------
FFT over 2,400,000 random values____________________ (sec): 1.23466666666667
Eigenvalues of a 640x640 random matrix______________ (sec): 1.15366666666667
Determinant of a 2500x2500 random matrix____________ (sec): 1.92633333333333
Cholesky decomposition of a 3000x3000 matrix________ (sec): 1.81366666666667
Inverse of a 1600x1600 random matrix________________ (sec): 1.96333333333333
--------------------------------------------
Trimmed geom. mean (2 extremes eliminated): 1.6278443559224
III. Programmation
------------------
3,500,000 Fibonacci numbers calculation (vector calc)(sec): 1.07533333333332
Creation of a 3000x3000 Hilbert matrix (matrix calc) (sec): 0.792999999999997
Grand common divisors of 400,000 pairs (recursion)__ (sec): 3.393
Creation of a 500x500 Toeplitz matrix (loops)_______ (sec): 1.20000000000001
Escoufier's method on a 45x45 matrix (mixed)________ (sec): 0.89500000000001
--------------------------------------------
Trimmed geom. mean (2 extremes eliminated): 1.04917789012305
Total time for all 15 tests_________________________ (sec): 22.6473333333333
Overall mean (sum of I, II and III trimmed means/3)_ (sec): 1.30868512419174
--- End of test ---
open source 2.11.1 32 bit
R version 2.11.1 (2010-05-31)
Copyright (C) 2010 The R Foundation for Statistical Computing
ISBN 3-900051-07-0
R is free software and comes with ABSOLUTELY NO WARRANTY.
You are welcome to redistribute it under certain conditions.
Type 'license()' or 'licence()' for distribution details.
Natural language support but running in an English locale
R is a collaborative project with many contributors.
Type 'contributors()' for more information and
'citation()' on how to cite R or R packages in publications.
Type 'demo()' for some demos, 'help()' for on-line help, or
'help.start()' for an HTML browser interface to help.
Type 'q()' to quit R.
[R.app GUI 1.34 (5589) i386-apple-darwin9.8.0]
[Workspace restored from /Users/jbreen/.RData]
trying URL 'http://watson.nci.nih.gov/cran_mirror/bin/macosx/leopard/contrib/2.11/SuppDists_1.1-8.tgz'
Content type 'application/x-gzip' length 412485 bytes (402 Kb)
opened URL
==================================================
downloaded 402 Kb
The downloaded packages are in
/var/folders/+s/+snLnQILHz4Kn8jMFYEERE++-4+/-Tmp-//RtmpX3gq6O/downloaded_packages
> source("/Users/jbreen/Desktop/R-benchmark-25.R")
Loading required package: Matrix
Loading required package: lattice
Attaching package: 'Matrix'
The following object(s) are masked from 'package:base':
det
Loading required package: SuppDists
R Benchmark 2.5
===============
Number of times each test is run__________________________: 3
I. Matrix calculation
---------------------
Creation, transp., deformation of a 2500x2500 matrix (sec): 1.19433333333333
2400x2400 normal distributed random matrix ^1000____ (sec): 1.122
Sorting of 7,000,000 random values__________________ (sec): 1.144
2800x2800 cross-product matrix (b = a' * a)_________ (sec): 2.25133333333333
Linear regr. over a 3000x3000 matrix (c = a \ b')___ (sec): 1.475
--------------------------------------------
Trimmed geom. mean (2 extremes eliminated): 1.26312946510292
II. Matrix functions
--------------------
FFT over 2,400,000 random values____________________ (sec): 1.40366666666667
Eigenvalues of a 640x640 random matrix______________ (sec): 1.28133333333333
Determinant of a 2500x2500 random matrix____________ (sec): 1.745
Cholesky decomposition of a 3000x3000 matrix________ (sec): 1.61
Inverse of a 1600x1600 random matrix________________ (sec): 1.50166666666667
--------------------------------------------
Trimmed geom. mean (2 extremes eliminated): 1.50275368328927
III. Programmation
------------------
3,500,000 Fibonacci numbers calculation (vector calc)(sec): 1.22666666666666
Creation of a 3000x3000 Hilbert matrix (matrix calc) (sec): 0.87133333333333
Grand common divisors of 400,000 pairs (recursion)__ (sec): 1.29
Creation of a 500x500 Toeplitz matrix (loops)_______ (sec): 1.33899999999999
Escoufier's method on a 45x45 matrix (mixed)________ (sec): 1.16800000000001
--------------------------------------------
Trimmed geom. mean (2 extremes eliminated): 1.22721231763175
Total time for all 15 tests_________________________ (sec): 20.6233333333333
Overall mean (sum of I, II and III trimmed means/3)_ (sec): 1.32561819861344
--- End of test ---